Weather anomalies

Climate change and temperature anomalies
If we wanted to study climate change, we can find data on the Combined Land-Surface Air and Sea-Surface Water Temperature Anomalies in the Northern Hemisphere at NASA’s Goddard Institute for Space Studies. The tabular data of temperature anomalies can be found here
To define temperature anomalies you need to have a reference, or base, period which NASA clearly states that it is the period between 1951-1980.
weather <-
read_csv("https://data.giss.nasa.gov/gistemp/tabledata_v4/NH.Ts+dSST.csv",
skip = 1,
na = "***")For each month and year, the dataframe shows the deviation of temperature from the normal (expected). Further the dataframe is in wide format.
Cleaning Data
tidyweather <- weather %>%
select(1:13) %>%
pivot_longer( cols = 2:13,
names_to = "Month",
values_to = "delta")
glimpse(tidyweather)## Rows: 1,716
## Columns: 3
## $ Year <dbl> 1880, 1880, 1880, 1880, 1880, 1880, 1880, 1880, 1880, 1880, 1880…
## $ Month <chr> "Jan", "Feb", "Mar", "Apr", "May", "Jun", "Jul", "Aug", "Sep", "…
## $ delta <dbl> -0.39, -0.53, -0.23, -0.30, -0.05, -0.18, -0.21, -0.25, -0.24, -…Plotting Information
Plotting data using a time-series scatterplot with a trendline.
tidyweather <- tidyweather %>%
mutate(date = ymd(paste(as.character(Year), Month, "1")),
month = month(date, label=TRUE),
year = year(date))
ggplot(tidyweather, aes(x=date, y = delta))+
geom_point()+
geom_smooth(color="red") +
theme_bw() +
labs (
title = "Weather Anomalies",
x = "Year",
y = "Temperature deviaton",
caption = "Source: https://data.giss.nasa.gov/gistemp/tabledata_v4/NH.Ts+dSST.txt"
) +
NULL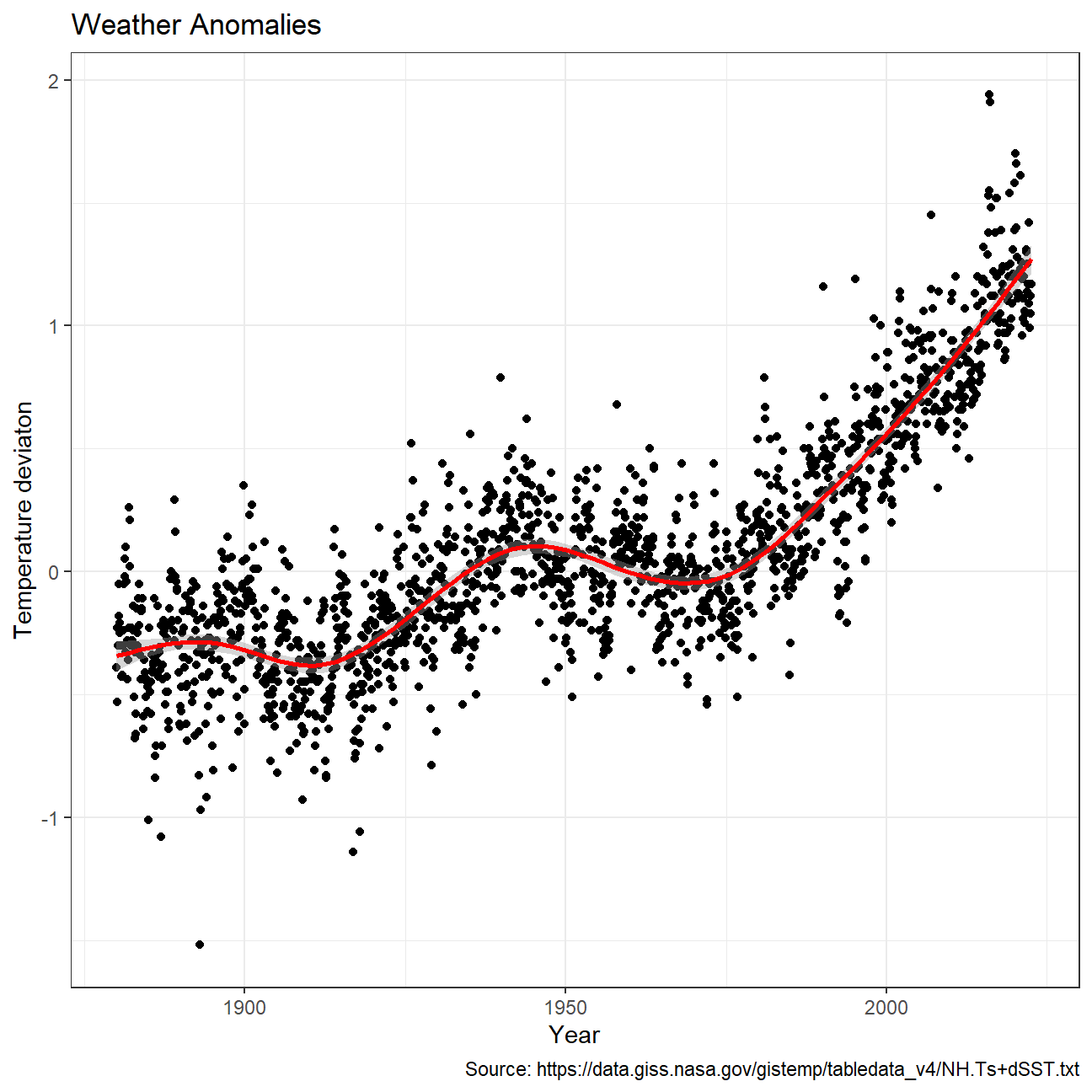
Producing a scatter plot showing the temperature anomalies by month.
ggplot(tidyweather, aes(x=date, y = delta))+
geom_point()+
geom_smooth(color="red") +
theme_bw() +
labs (
title = "Weather Anomalies",
x = "Year",
y = "Temperature deviaton",
caption = "Source: https://data.giss.nasa.gov/gistemp/tabledata_v4/NH.Ts+dSST.txt"
) +
facet_wrap(~month) +
NULL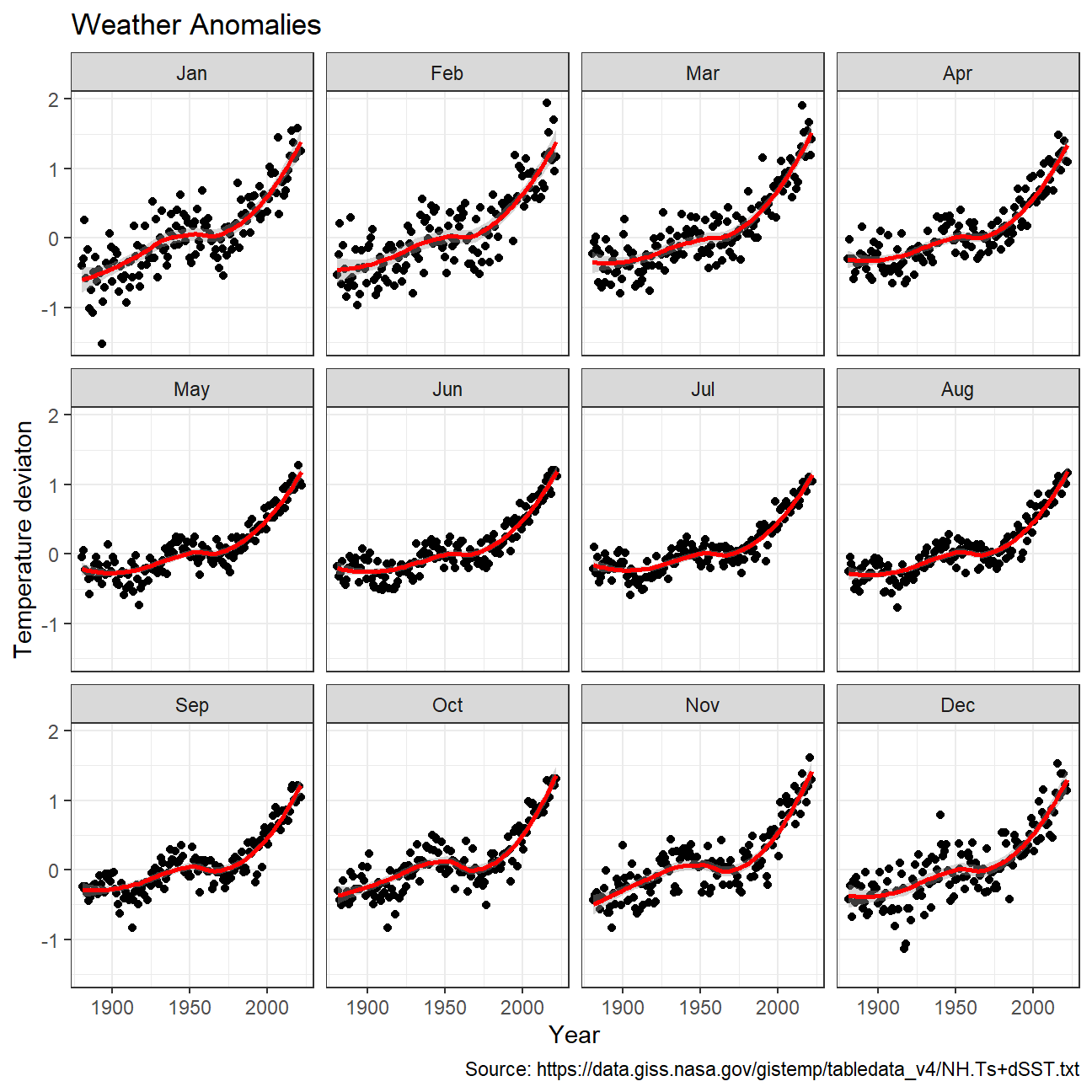
Grouping data into different time periods to study historical data.
comparison <- tidyweather %>%
filter(Year>= 1881) %>% #remove years prior to 1881
#create new variable 'interval', and assign values based on criteria below:
mutate(interval = case_when(
Year %in% c(1881:1920) ~ "1881-1920",
Year %in% c(1921:1950) ~ "1921-1950",
Year %in% c(1951:1980) ~ "1951-1980",
Year %in% c(1981:2010) ~ "1981-2010",
TRUE ~ "2011-present"
))Creating a density plot to study the distribution of monthly deviations grouped by the different time periods.
ggplot(data = comparison, aes(delta)) +
geom_density(aes(fill = interval), alpha = 1/4) +
labs(title = "Distribution of Monthly Temperature Anomalies in Time Intervals",
x = "Monthly Temperature Anomaly",
y = "Density",
caption = "Source: https://data.giss.nasa.gov/gistemp/tabledata_v4/NH.Ts+dSST.txt") +
facet_wrap(~ interval, ncol = 1) +
theme_bw() +
theme(legend.position = "none") +
NULL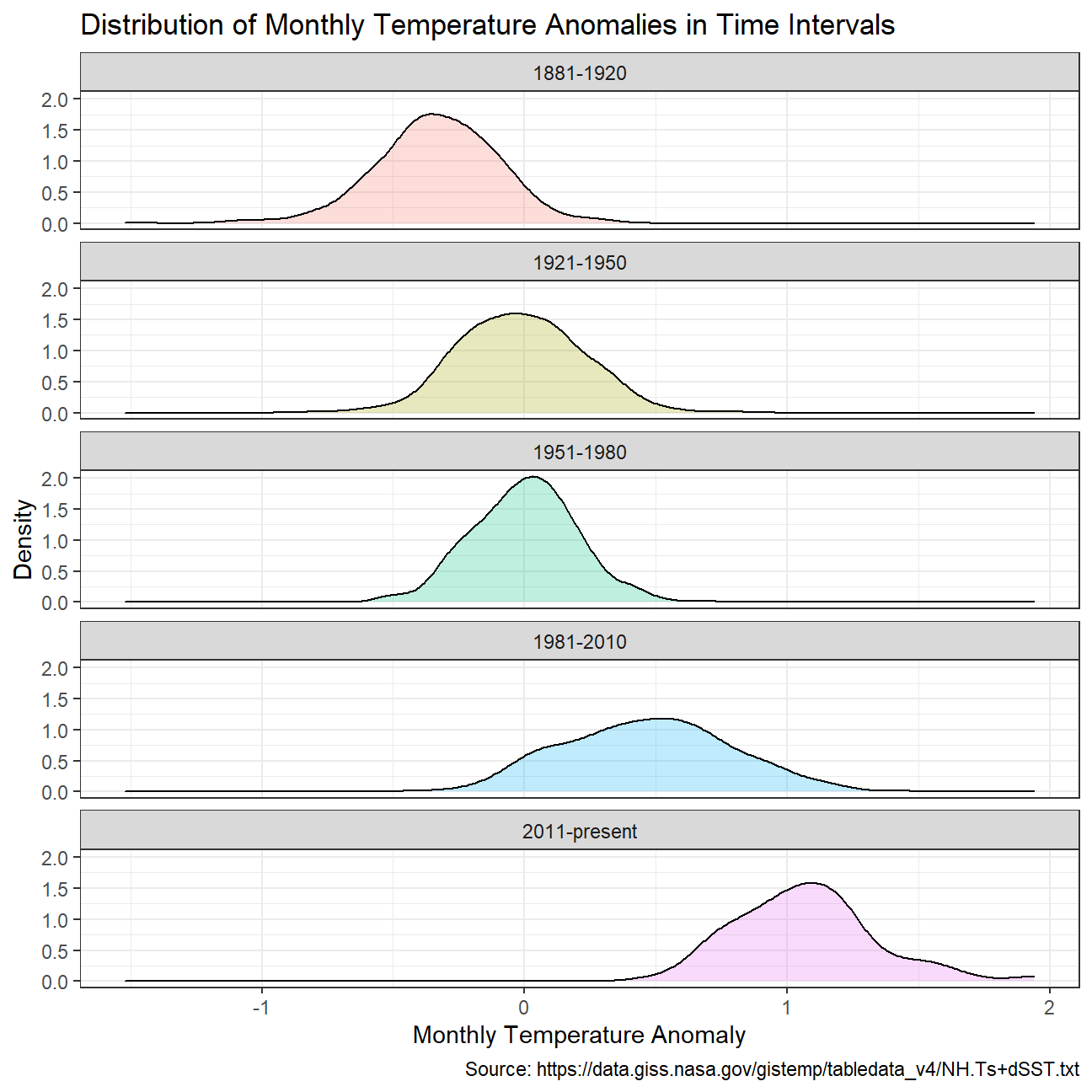
Calculating average annual anomalies.
#creating yearly averages
average_annual_anomaly <- tidyweather %>%
group_by(Year) %>% #grouping data by Year
# creating summaries for mean delta
summarise(mean_delta = mean(delta, na.rm=TRUE))
#plotting the data:
ggplot(average_annual_anomaly,
aes (x = Year,
y = mean_delta)) +
geom_point() +
theme_bw() +
geom_smooth(method = "loess") +
labs(title = "Average annual anomalies by year",
y = "Average annual anomalies",
caption = "Source: https://data.giss.nasa.gov/gistemp/tabledata_v4/NH.Ts+dSST.txt") +
NULL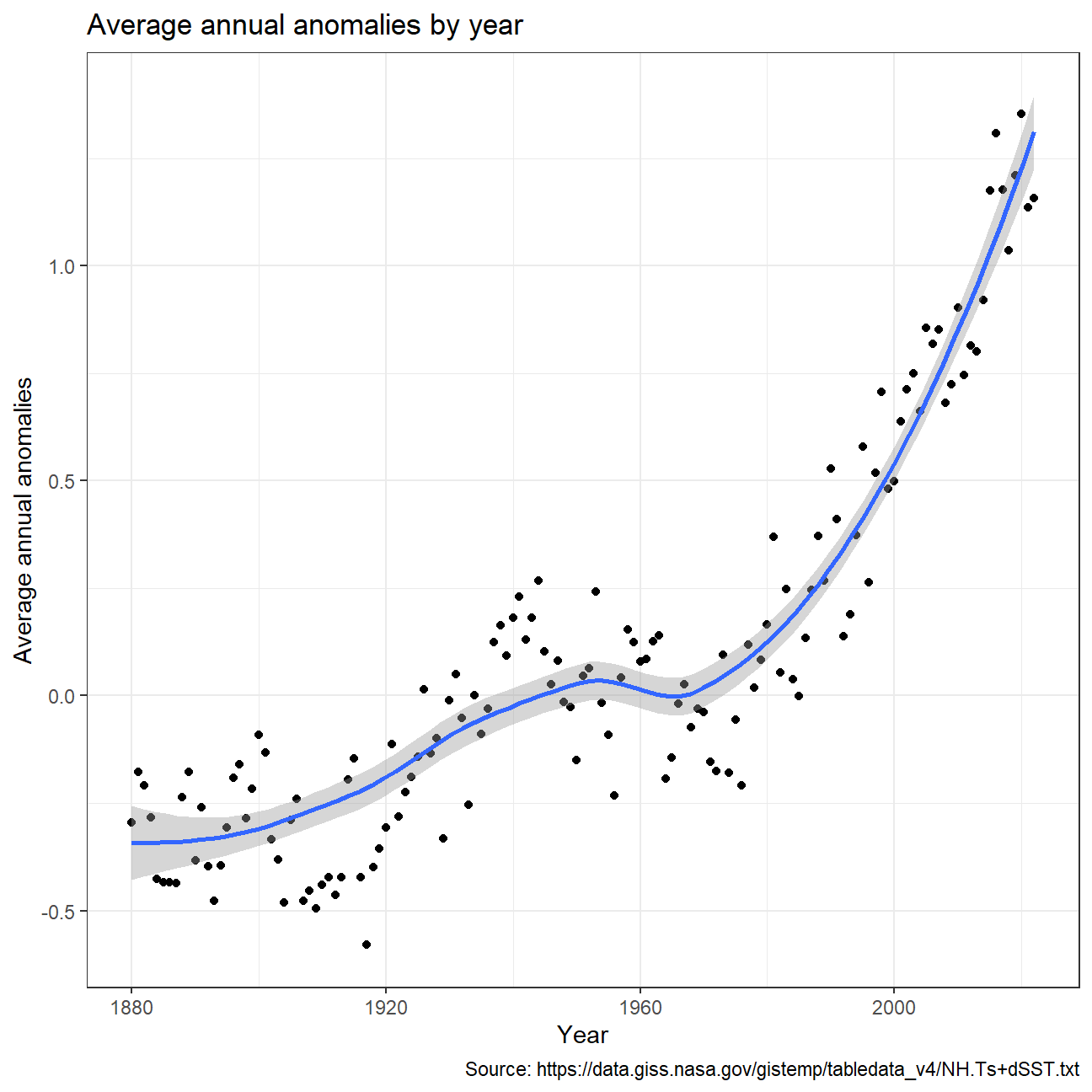
Confidence Interval for delta
NASA points out on their website that
A one-degree global change is significant because it takes a vast amount of heat to warm all the oceans, atmosphere, and land by that much. In the past, a one- to two-degree drop was all it took to plunge the Earth into the Little Ice Age.
Construction of a confidence interval for the average annual delta since
2011, both using a formula and using a bootstrap simulation with the
infer package.
formula_ci <- comparison %>%
filter(interval == "2011-present") %>% # choose the interval 2011-present
filter(!delta == "NA") %>% # drop NA observations in delta
summarise(count = n(),
t = qt(0.975, count-1), # use qt with probability and degrees of freedom
mean = mean(delta), # calculate mean
sd = sd(delta), # calculate sd
se = sd(delta)/sqrt(count), # calculate se
margin = t * se, # calculate margin of error
lower = mean - margin, # calculate lower bound
upper = mean + margin #calculate upper bound
)
#print out formula_CI
formula_ci## # A tibble: 1 × 8
## count t mean sd se margin lower upper
## <int> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 140 1.98 1.07 0.265 0.0224 0.0443 1.02 1.11library(infer)
set.seed(1234)
boot_ci <- comparison %>%
filter(interval == "2011-present") %>% # choose the interval 2011-present
filter(!delta == "NA") %>% # drop NA observations in delta
specify(response = delta) %>% # specify the variable of interest
generate(reps = 1000, type = "bootstrap") %>% # extract 1000 bootstrap samples
calculate(stat = "mean") %>% # calculate sample means from each bootstrap sample
get_confidence_interval(level = 0.95, type = "percentile") # calculate confidence interval of this analysis
# Display confidence interval
boot_ci## # A tibble: 1 × 2
## lower_ci upper_ci
## <dbl> <dbl>
## 1 1.02 1.11Two different methods of constructing 95% confidence interval were used in this example. First was based on filtering appropriate interval and calculating confidence interval using summary statistics. Second involved ‘infer’ package, which allowed to use bootstrap method and produced the confidence intervals without any additional summary statistics. According to the summary calculations the average annual anomalies since 2011 already exceeded 1 degree, even when 95% confidence interval is taken into account. Therefore, it is highly likely that anomalies will become even more frequent and significant than before.